Topic 7.3: Translation
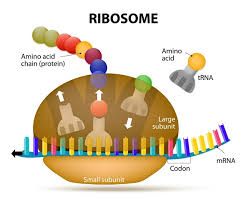 image from News Medical
image from News Medical
In the Translation Unit we will extend our knowledge on Topic 2.7 DNA Replication, Transcription and Translation.
Translation is the second part of the central dogma of molecular biology, describing how the genetic code is used to make amino acid chains. This unit will explore the mechanics involved in polypeptide synthesis. You will learn the three major steps of translation as tRNA, mRNA, and ribosomes form the polypeptide. We will also explore the structural formation of polypeptides after translation
This unit will last 3 school days
Translation is the second part of the central dogma of molecular biology, describing how the genetic code is used to make amino acid chains. This unit will explore the mechanics involved in polypeptide synthesis. You will learn the three major steps of translation as tRNA, mRNA, and ribosomes form the polypeptide. We will also explore the structural formation of polypeptides after translation
This unit will last 3 school days
Essential idea:
- Information transferred from DNA to mRNA is translated into an amino acid sequence.
Nature of science:
- Developments in scientific research follow improvements in computing—the use of computers has enabled scientists to make advances in bioinformatics applications such as locating genes within genomes and identifying conserved sequences. (3.7)
Understandings:
7.3 U 1 Initiation of translation involves assembly of the components that carry out the process. (Oxford Biology Course Companion page 364).
- Outline the process of translation initiation.
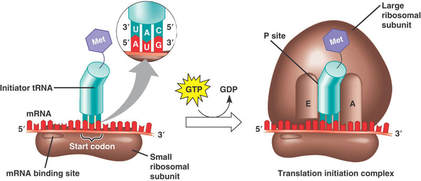
The first stage of translation involves the assembly of the three components that carry out the process (mRNA, tRNA, ribosome)
Start codon:
Initiation:
Start codon:
- the mRNA triplet codon AUG is universally the start codon used to mark the beginning of the coding sequence of a gene;
- the tRNA with the anticodon UAC, and carrying the amino acid methionine, is always the first tRNA to enter the P-site during translation
Initiation:
- 5’ end of mRNA binds to the small subunit of the ribosome
- initial mRNA codon = AUG = start codon
- tRNA with anticodon: UAC binds to mRNA AUG codon by complementary base pairing, methionine attached to tRNA 3’ terminal
- large ribosomal subunit binds, completing ribosomal structure, and producing two ribosomal binds sites: P site & A site
7.3 U 2 Synthesis of the polypeptide involves a repeated cycle of events. (Oxford Biology Course Companion page 365).
- Outline the process of translation elongation, including codon recognition, bond formation and translocation.
- State the direction of movement of the ribosome along the mRNA molecule.
Elongation
Translocation
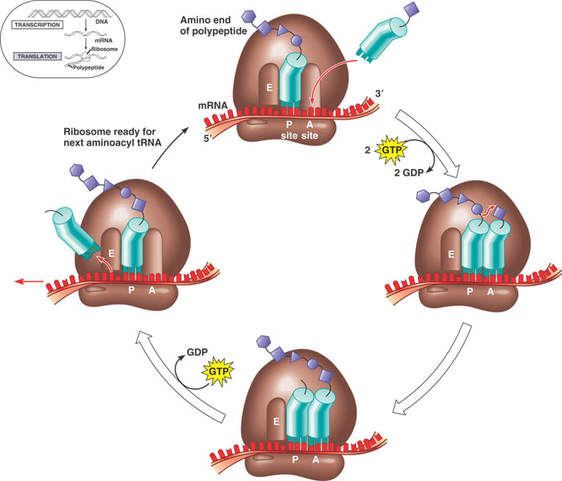
- While the first tRNA is still attached, a second tRNA attaches to the mRNA at the A site on the ribosome, carrying the amino acid that corresponds to the mRNA codon.
- The methionine amino acid at the P site binds to the amino acid carried by the second tRNA located at the A site.
- The two amino acids are joined together through a condensation reaction that creates a peptide bond between the two amino acids.
Translocation
- The ribosome moves along the mRNA one codon shifting the tRNA that was attached to methionine to the E site.
- The tRNA is released back into the cytoplasm from the E site, allowing it to pick up another amino acid (methionine) to build another polypeptide.
- Another tRNA moves into the empty A site bringing the next amino acid corresponding to the mRNA codon.
- Again, the amino acid is attached to the polypeptide forming a peptide bond, the ribosome slides across one codon and tRNA at the P site moves into the E site releasing it back into the cytoplasm.
- The ribosome continues to move along the mRNA adding amino acids to the polypeptide chain.
- This process continues until a stop codon is reached.
7.3 U 3 Disassembly of the components follows termination of translation. (Oxford Biology Course Companion page 365).
- Outline the process of translation termination, including the role of the stop codon.
Termination:
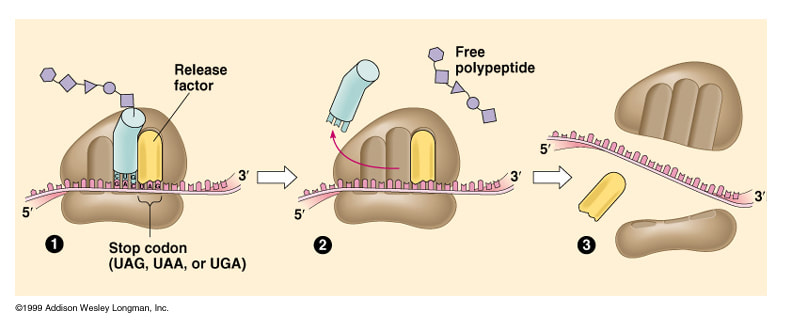
- when ribosomal A-site reaches a stop codon, no tRNA has a complementary anticodon
- there are three stop codons in the genetic code; none of these have a corresponding tRNA
- when a ribosome encounters a stop codon, a release factor binds to the stop codon
- release factor protein binds to ribosomal A-site stop codon
- polypeptide and mRNA are released
- large and small ribosomal subunits separate
- Ribosome reaches the STOP codon. Release factor attaches
- tRNA released to find another amino acid. Polypeptide released
- Components of the ribosomoe break apart. All are used again
7.3 U 4 Free ribosomes synthesize proteins for use primarily within the cell. (Oxford Biology Course Companion page 366).
- State the difference between free and bound ribosomes.
- List destinations of proteins synthesized on free ribosomes.
|
Ribosomes are composed of rRNA and protein.
Free ribosomes are located in the cytoplasm, and the bound ribosomes are attached to the endoplasmic reticulum. Free ribosomes produce proteins for the cell, while bound ribosomes produce proteins that are transported out of the cell. Free ribosomes in the cytoplasm synthesize proteins that will be used inside the cell in the cytoplasm, mitochondria and chloroplasts (in autotrophs). Free ribosomes synthesize proteins for use primarily within the cell itself that bound ribosomes synthesize proteins primarily for secretion or for lysosomes |
7.3 U 5 Bound ribosomes synthesize proteins primarily for secretion or for use in lysosomes. (Oxford Biology Course Companion page 366).
- List destinations of proteins synthesized on bound ribosomes.
- Outline how a ribosome becomes bound to the endoplasmic reticulum.
Ribosomes attached to ER create proteins that are secreted from the cell by exocytosis or are used in lysosomes.
Proteins perform many functions within specific compartments of the cell or in other parts of the body after they are secreted out of the cell
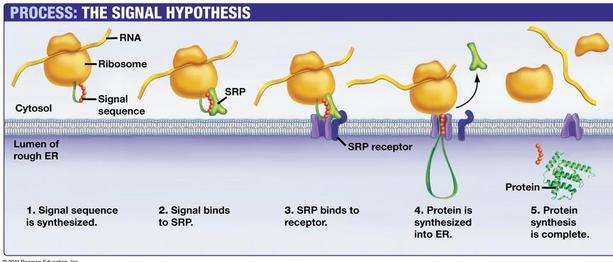
Proteins that are destined to be used in lysosomes, ER, Golgi Apparatus, the plasma membrane or secreted by the cell are made by ribosomes bound by the endoplasmic reticulum. Ribosomes that become bound to the ER are directed here by a signal sequence that is part of that specific polypeptide. This signal sequence on the polypeptide binds to a signal recognition protein (SRP). The SRP guides the polypeptide and ribosome to the ER where it binds to an SRP receptor. Translation can now continue and the polypeptide is deposited into the lumen of the ER as its created for transportation to the correct location
Proteins perform many functions within specific compartments of the cell or in other parts of the body after they are secreted out of the cell
Proteins that are destined to be used in lysosomes, ER, Golgi Apparatus, the plasma membrane or secreted by the cell are made by ribosomes bound by the endoplasmic reticulum. Ribosomes that become bound to the ER are directed here by a signal sequence that is part of that specific polypeptide. This signal sequence on the polypeptide binds to a signal recognition protein (SRP). The SRP guides the polypeptide and ribosome to the ER where it binds to an SRP receptor. Translation can now continue and the polypeptide is deposited into the lumen of the ER as its created for transportation to the correct location
7.3 U 6 Translation can occur immediately after transcription in prokaryotes due to the absence of a nuclear membrane. (Oxford Biology Course Companion page 365).
- Compare the timing and location of transcription and translation between prokaryotes and eukaryotes.
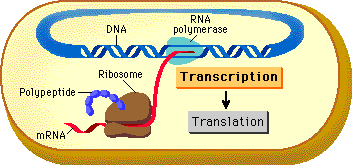
Since prokaryotic DNA is not compartmentalized into a nucleus, once transcription begins creating a strand of mRNA, translation can begin immediately as the mRNA strand is created. In eukaryotes, the completed mRNA has to be transported from the nucleus, through the nuclear pore to the ribosome on the ER or in the cytosol
7.3 U 7 The sequence and number of amino acids in the polypeptide is the primary structure. (Oxford Biology Course Companion page 366).
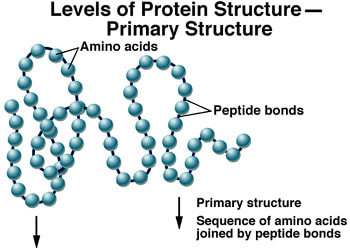
- Describe the primary structure of a protein, including the type of bonding involved.
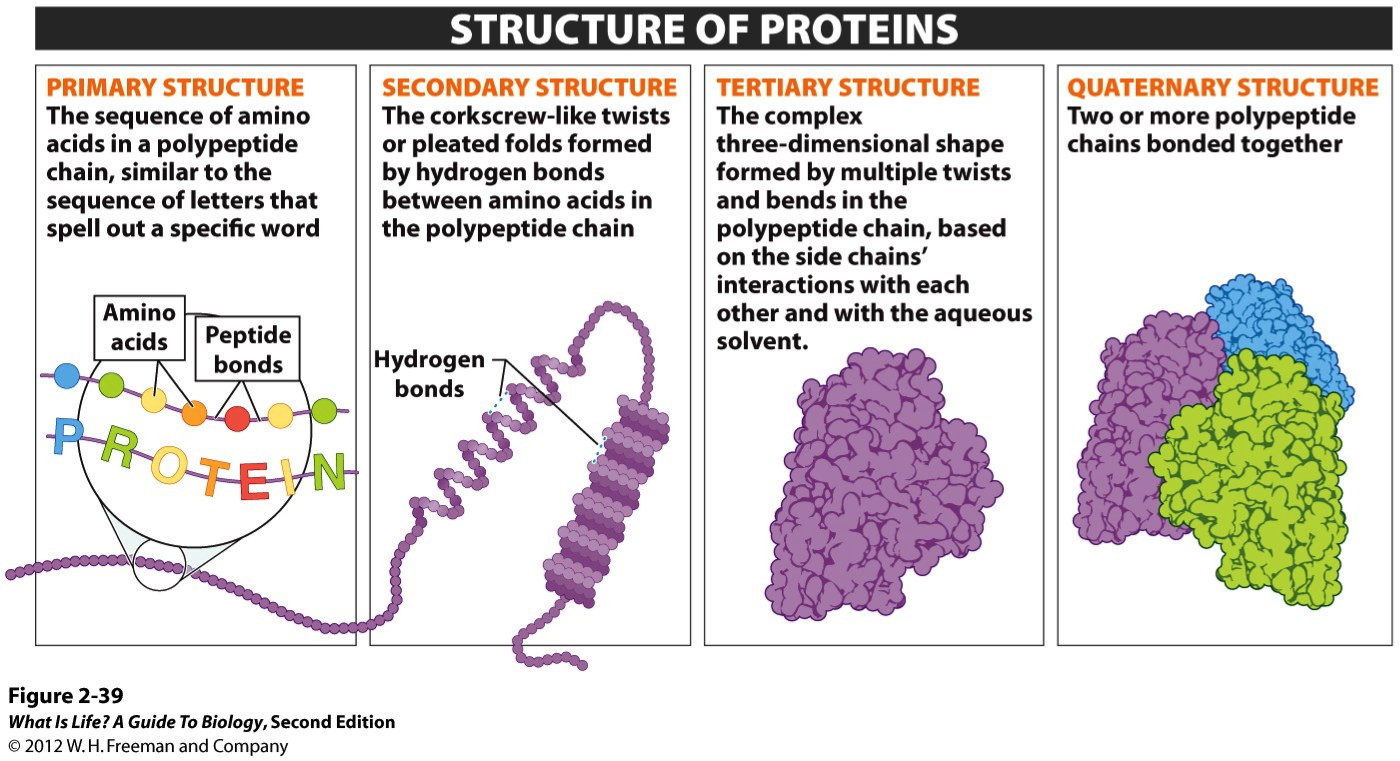
The simplest level of protein structure, primary structure, is simply the sequence of amino acids in a polypeptide chain. Each chain has its own set of amino acids, assembled in a particular order. The sequence can consist of any of the 21 amino acids and is coded for by a gene in the DNA. The sequence of amino acids can determine the shape of the other three levels of protein structure.
7.3 U 8 The secondary structure is the formation of alpha helices and beta pleated sheets stabilized by hydrogen bonding. (Oxford Biology Course Companion page 120).
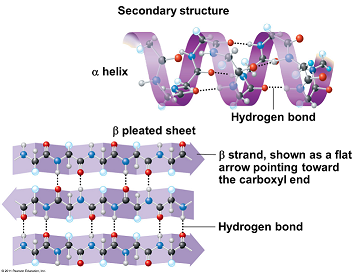
- Describe the secondary structure of a protein, including the type of bonding involved.
- Identify the alpha-helix and beta-pleated sheet in images of protein structure.
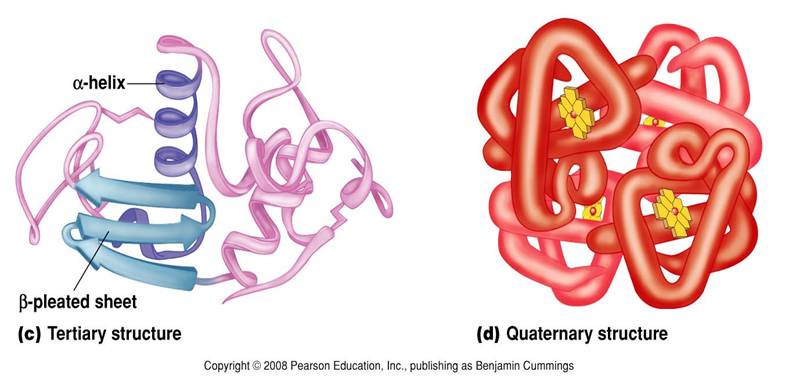
Secondary structure is created when the hydrogen bonds are formed between the main peptide groups in amino acids (not the R groups) in the polypeptide. The two main structures formed are the alpha helix and the beta pleated sheets which are locally defined, which means there can be many different secondary structures present in a single protein molecule. Proteins with secondary structure are generally structural in nature like the secondary structure in silk.
Weak hydrogen bonds between amino and carboxyl groups and different amino acids form at regular intervals, creating a regular structure (not from interactions between variable R- groups)
Weak hydrogen bonds between amino and carboxyl groups and different amino acids form at regular intervals, creating a regular structure (not from interactions between variable R- groups)
- alpha helix = coiling into a helix
- beta pleating = a folded sheet as polypeptide folds back on itself
- not all of a polypeptide forms secondary structure in most proteins
7.3 U 9 The tertiary structure is the further folding of the polypeptide stabilized by interactions between R groups.
- Describe the tertiary structure of a protein, including the types of R group interactions involved.
- Explain how the chemical characteristics of R groups in the polypeptide chain affect protein folding.
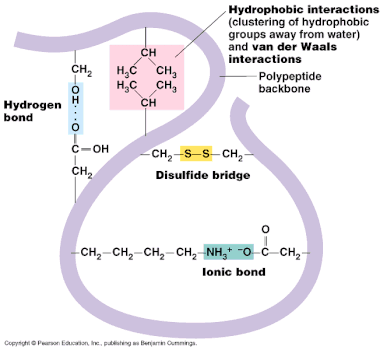
Tertiary structure is the third level of protein organization. Tertiary structure develops a three dimensional shape because of the interactions that occur between the R groups.
- Disulphide bridges form between sulphur atoms on the R-groups from the amino acid cysteine.
- Hydrogen bonds form between polar R-groups and other polar R-groups.
- Ionic Bonds are also formed between some R-groups
- Hydrophobic R-groups turn inwards away from water and hydrophilic R-groups face outwards towards the water.
- Protein is globular in nature.
- Interactions between variable R- groups forming:
- hydrophobic interactions between nonpolar amino acids
- hydrogen bonds between polar amino acids
- ionic bonds between ionic amino acids
- covalent bonds between sulfur containing amino acids
- Producing the three-dimensional folded structure of most proteins
7.3 U 10 The quaternary structure exists in proteins with more than one polypeptide chain.
- Outline the quaternary structure of protein folding.
- Describe the structure of a conjugated protein, including the prosthetic group.
Protein consisting of more than one polypeptide chain. These chains can consist of quaternary structures. Quaternary structure exists in many proteins such as hemoglobin (4 polypeptide chains).
Proteins with prosthetic groups are called conjugated proteins. Aggregations of polypeptides form interactions between more than one polypeptide
- Hemoglobin also contains a prosthetic group (non-polypeptide structure). The prosthetic group in hemoglobin is iron. Hemoglobin contains 4 iron molecules.
Proteins with prosthetic groups are called conjugated proteins. Aggregations of polypeptides form interactions between more than one polypeptide
- polypeptides + non-proteinaceous molecules, such as:
- metals, e.g. iron in hemoglobin
- vitamins as enzyme cofactors
- nucleic acids as in ribosomes
- carbohydrates in glycoproteins
- lipids in lipoproteins
Application
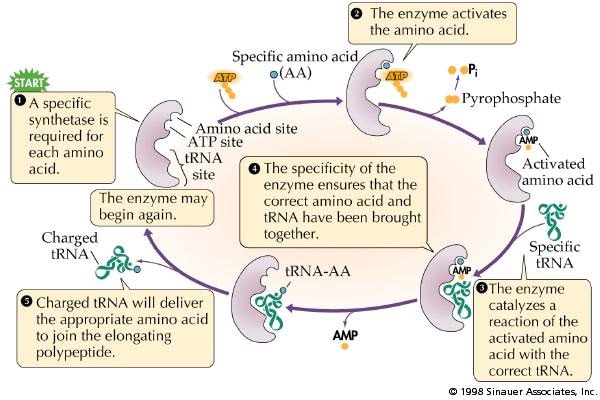
7.3 A 1 tRNA-activating enzymes illustrate enzyme–substrate specificity and the role of phosphorylation. (Oxford Biology Course Companion page 364).
- State the role of the tRNA activating enzymes.
- Outline the process of attaching an amino acid to tRNA by the tRNA activating enzyme.
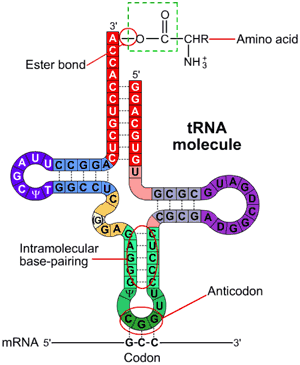
Each tRNA binds with a specific amino acid in the cytoplasm in a reaction catalyzed by a specific tRNA-activating enzyme (21 specific enzymes for the 21 different amino acids). Each specific amino acid binds covalently to the 3'- terminal nucleotide (CCA) at the end of the tRNA molecule. The binding of the specific amino acid to the tRNA requires energy from ATP.
There are many different types of tRNA in a cell.All have tRNA molecules have:
There are many different types of tRNA in a cell.All have tRNA molecules have:
- a triplet of bases called the anticodon, found in the anticodon loop of 7 nucleotides
- two other loops
- a CCA base sequence at the 3’ terminal, which forms a site for attaching an amino acid d) sections that become double-stranded by base pairing
Skills
7.3.S1 The use of molecular visualization software to analyze the structure of eukaryotic ribosomes and tRNA molecules (Oxford Biology Course Companion page 362).
- Describe the structure of the ribosomes, including the small and large subunits and the names and roles of the tRNA binding sites.
- Use molecular visualization software to view and identify the small and large subunit and tRNA binding sites of the ribosome.
- Outline the structure of tRNA molecules.
- Use molecular visualization software to view and identify the anticodon and amino acid binding site of a tRNA.
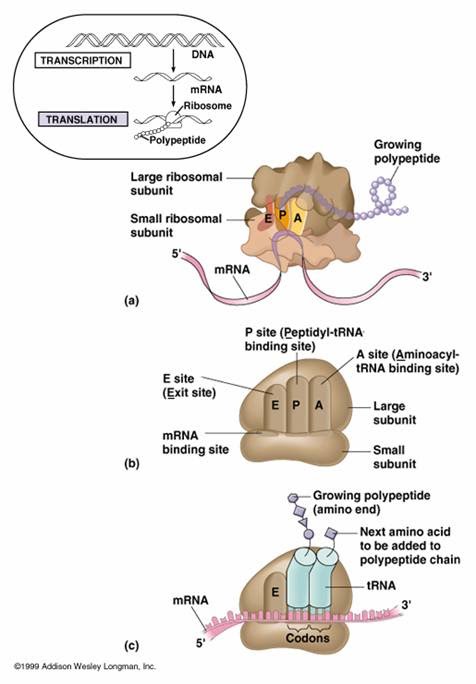
Ribosomes are organelles made from protein and RNA that catalyze the assembly of amino acids into polypeptides during translation. Ribosomes consist of two subunits; a large subunit and a small sub unit. The large sub-unit consists of 3 binding sites for tRNA molecules called the A site (The A site binds an aminoacyl-tRNA (tRNA bound to an amino acid); the P site binds a peptidyl-tRNA (tRNA bound to the peptide being synthesized); and the E site binds a free tRNA (no amino acid attached) before it exits the ribosome. The order of the sites in the ribosomes is E P A.
There is a binding site for the mRNA on the surface of the ribosome The small subunit of the ribosome contains the binding site for the mRNA strand. The space between the two subunits of the ribosome is where the polypeptide is assembled during translation.
There is a binding site for the mRNA on the surface of the ribosome The small subunit of the ribosome contains the binding site for the mRNA strand. The space between the two subunits of the ribosome is where the polypeptide is assembled during translation.
7.3 S 2 Identification of polysomes in electron micrographs of prokaryotes and eukaryotes. (Oxford Biology Course Companion page 366).
- Outline the structure of a polysome.
- Identify the beginning of an mRNA strand in a micrograph of polysomes.
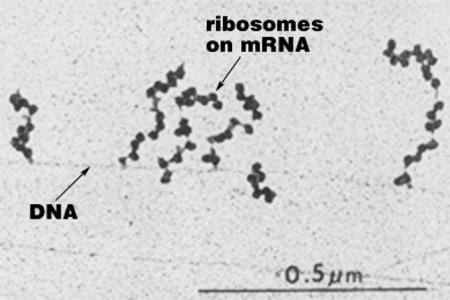
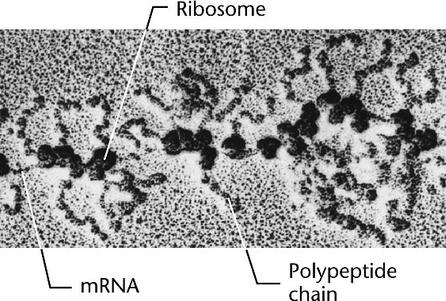
Polysomes are several ribosomes used to translate an mRNA at the same time, forming what is called a polysome. More than one ribosome can translate an mRNA at one time, making it possible to produce many polypeptides simultaneously from a single mRNA.
In prokaryotes, several ribosomes can attach themselves to the growing mRNA chains to form a polysome while the mRNA chains are still attached to the DNA (left)
In eukaryotes, the mRNA detaches from the DNA and is then transported through pores in the nuclear envelope to the ribosomes in the cytoplasm. Once in the cytosol, eukaryote mRNA can also form polysomes (right)
In prokaryotes, several ribosomes can attach themselves to the growing mRNA chains to form a polysome while the mRNA chains are still attached to the DNA (left)
In eukaryotes, the mRNA detaches from the DNA and is then transported through pores in the nuclear envelope to the ribosomes in the cytoplasm. Once in the cytosol, eukaryote mRNA can also form polysomes (right)
Key Terms
|
amino acid sequence
codon recognition stop codon protein synthesis endoplasmic reticulum tertiary structure tRNA activating enzymes phosphorylation |
initiation
polypeptide free ribosomes alpha helices protein folding prosthetic group bioinformatics |
translationribosome
bound ribosomes primary structure hydrogen bonding conjugated protein genome analysis |
elongation
mRNA lysosomes secondary structure beta pleated sheets quaternary structure enzyme-substrate |
initiation
translation termination R groups binding sites anticodons polysome |
PowerPoint and Notes on Topic 7.3 by Chris Payne
Your browser does not support viewing this document. Click here to download the document.
Your browser does not support viewing this document. Click here to download the document.
Correct use of terminology is a key skill in Biology. It is essential to use key terms correctly when communicating your understanding, particularly in assessments. Use the quizlet flashcards or other tools such as learn, scatter, space race, speller and test to help you master the vocabulary.
Useful Links
Translation: DNA to mRNA to Protein
Stop! Codon time!
Nucleic Acids to Amino Acids: DNA Specifies Protein
DNA to Protein animation
Modeling Translation animation
In The News
Protein 'intentionally' terminates own synthesis by destabilizing synthesis machinery -- the ribosome - ScienceDaily, Nov 2017
Unknown Protein Structures Predicted - The Scientists, Jan 2017
Translation: DNA to mRNA to Protein
Stop! Codon time!
Nucleic Acids to Amino Acids: DNA Specifies Protein
DNA to Protein animation
Modeling Translation animation
In The News
Protein 'intentionally' terminates own synthesis by destabilizing synthesis machinery -- the ribosome - ScienceDaily, Nov 2017
Unknown Protein Structures Predicted - The Scientists, Jan 2017
Video Clips
Transcription in eukaryotes Animation
Translation process for protein synthesis.
tRNA
Everyone has heard of proteins. What are they on the molecular level? They're polymers of amino acids, of course. They make up most of your body, so we have to understand their structure very well! Check this out to learn the hierarchy of protein structure so that we can later learn all about what different types of proteins can do.